Season 1
General info on decoding MOM6 sessions
- The early sessions of these “Decoding MOM6” segments will talk through the tools that might be useful to navigate the MOM6 code
- The later sessions will dive deeper into the code itself
How to use github's search functionality to find code: MOM6 as an example.
Presenter: @dougiesquire
Searching the MOM6 code repository in the ACCESS-NRI GitHub org. Contains all of the MOM6 Fortran code used in ACCESS-OM3
Quick summary of GitHub search functionality
Pros:
- Easy and convenient
- Allows you to see how parts of the model work
Cons:
- Can only search on the default branch of the repo.
- GitHub doesn't actually index everything.
More detailed notes
- Why use the GitHub search bar?
- See how a specific piece of code is written - how is it implemented in MOM6? Need to search through the code
- Dougie’s screen: Looking at MOM6 fork on ACCESS-NRI GitHub org
- Can click on search at top or click
/to start a search - Eg search
global- Will return all files in this repo with “Global” (not case sensitive by default)
- Get the summary of the files, and can click to expand more lines of code
- Can click on the line itself and it will take you to that line in the file
- Can search on multiple terms
- Eg
Global Mean- searches all files with global or mean - Eg
“Global Mean”- search for specific phrase - Eg
“Global mean ocean salinity”- can search directly for this diagnostic- Finds where it’s registered, and can backtrack to find the code
- Eg
- Can search on patterns (regex) rather than words
- Eg search
Globalbut only at specific path:Global path:src/core
- Eg search
- Fast, powerful, easy to use
- Can share searches with people (can’t do this if you are, eg, using “grep” on Gadi terminal)
- Can click on search at top or click
- Gotchas
- Can only search on default branch
- GitHub doesn’t index everything, and opaque which code is indexed
- Eg doesn’t index vendor code, but not clear what that means
- If you search something and it seems weird that nothing is found, then this might be the main issue
Questions from the audience
- Q: Can it organize the search in order of the subroutines called?
- MOM6 docs does include schematics on subroutines that are called from specific file
- But may not be generated anymore, so may be out of date
- MOM6 docs does include schematics on subroutines that are called from specific file
- Q: Can searches be case sensitive?
- Yes, you can use regex:
/(?-i)Global/ - By default they aren’t case sensitive
- Yes, you can use regex:
- Q: What is regex?
- Language for pattern matching
- Stands for regular expression
- Lots of resources online
- Just be sure to use the correct version of regex, as there are different types
- Can click “Search syntax tips” - link at bottom of search popup in GitHub - provides quick help for lots of common regex uses
- Q: OM3 configurations are all on a branch - does that mean we’ve eliminated the ability to search through OM3 configurations?
- Yes, we can’t search OM3 configurations at the moment using this tool, because there is almost nothing on the default branch. This is definitely a limitation.
- Some parameter documentation has been moved to main branch specifically to be able to search on them
- There are other ways to search through other branches, but no examples were given
- Lots of people would like GitHub to add functionality to search on a branch other than the default, but not the case currently
More resources
Understanding GitHub Code Search syntax
Lessons learned from MOM6 code development
Presenter: @jbisits (11/12/2025).
Tip
Take-home message: when starting development, take care with which version of the MOM6 code repository you fork from.
Background
When @jbisits started work on MOM6, he went to google and found the MOM6 codebase and then made a fork of mom-ocean/MOM6 (the central consortium repository). Actually if one is looking to contribute to the MOM6 codebase from Australia, there is an access-nri/MOM6 fork. They are different forks with different branches and code bases!
How do I find out which version of MOM6 I am currently using?
Go to the OM3 configuration that you are using example and find the config.yaml following lines:
Note these lines:
modules:
use:
- /g/data/vk83/modules
load:
- access-om3/2025.08.001
Specifically note that this line access-om3/2025.08.001 highlights the Spack bundle package and git tag that is being used for MOM6 code. One can then match this tag name 2025.08.001 from the model deployment repository, this link lists all the tags, and here is the tag we are looking for: 2025.08.001 in this case.
Further information:
- MOM6 fork management for being a development node;
- OM3 build system and deployment;
- NOAA-GFDL MOM6 development guide;
- MOM6 development presentation by @marshallward.
How to find code that corresponds to a particular executable
ACCESS-NRI executables
Presenter: @jbisits.
Scope: where and how to find which model components and source code are used in an OM3 configuration that are built with Spack (this applies to all ACCESS-NRI models).
Through ACCESS-NRI release database (friendly)
The easiest way to find a related version is to use the release database. For example, suppose we are interested in model version 2025.08.001 (listed in a config.yaml file in an access-om3-configs branch or directory if you have already cloned an experiment).
We can find this model version here. Then clicking on any of the github icons for the relevant component takes you to the version of the code that was used in that release.
Directly through GitHub (more complicated)
For example looking at ACCESS-OM3, we start by looking at the configuration repository: ACCESS-NRI/access-om3-configs. Note that the main branch does not contain configurations -- you have to look at the branches.
You may wish to know what are the model components being used in a particular release? Looking at the release: release-MC_25km_jra_iaf-1.0-beta we can browse the repository at the relevant tag here.
Then in the config.yaml, see the model software version:
modules:
use:
- /g/data/vk83/modules
load:
- access-om3/2025.08.001
access-om3/2025.08.001 refers to this repository with this at the tag: 2025.08.001. To find the tags one goes to the repository --> Releases --> Tags. Here is the related access-om3 release:
https://github.com/ACCESS-NRI/ACCESS-OM3/releases/tag/2025.08.001
Browsing the related spack.yaml we can see the versions of the components in the build, for example for CICE and MOM6 we have:
access-cice:
require:
- '@CICE6.6.1-0'
- io_type=PIO
- 'fflags="-march=sapphirerapids -mtune=sapphirerapids -unroll"'
- 'cflags="-march=sapphirerapids -mtune=sapphirerapids -unroll"'
- 'cxxflags="-march=sapphirerapids -mtune=sapphirerapids -unroll"'
access-mom6:
require:
- '@2025.07.000'
- 'fflags="-march=sapphirerapids -mtune=sapphirerapids -unroll"'
- 'cflags="-march=sapphirerapids -mtune=sapphirerapids -unroll"'
- 'cxxflags="-march=sapphirerapids -mtune=sapphirerapids -unroll"'
Like before, we go to the related ACCESS-NRI repositories and find the related tags, in this instance they are:
- access-CICE;
- access-mom6.
Note that the above two repositories are forks of the upstream repositories (CICE and MOM6)
For more information and other related steps: - https://access-om3-configs.access-hive.org.au/infrastructure/Building/ - https://docs.access-hive.org.au/getting_started/spack/ - https://docs.access-hive.org.au/models/build_a_model/build_source_code/ - https://docs.access-hive.org.au/models/build_a_model/create_a_prerelease/ (not shown in this tutorial but can be very simple)
COSIMA executables (e.g. mom6-panan)
Presenter: @angus-g.
Scope: where and how to find which model components and source code are used in a COSIMA configuration built with ninja.
COSIMA made executables use a different build system and way of tracking provenance. Here's an example using the mom6-panan. Going to the relevant line in the config.yaml we have: exe: /g/data/ik11/inputs/mom6/bin/symmetric_FMS2-e7d09b7
If one then goes on Gadi to the related folder, one finds:
/g/data/ik11/inputs/mom6/bin/manifest.yaml
Looking inside the manifest file we see a series of executables with information related to the major components (MOM6, SIS2, FMS) with the git hash used. The compiler flags are also included but not the more minor components. Below is an example:
exe: "symmetric_FMS2-e7d09b7"
build-date: 2022-01-20
sha256: "e7d09b7687fba4cf0693adf3c1a5f653dee312df996b99da384c64c279a53eef"
git:
- component: MOM6
ref: "93968bd8ec1a993c705846ec91c4cf33b86bc4fb"
- component: SIS2
ref: "d18605eacdd2c92b8262458fcad107c546dd080f"
- component: FMS
ref: "2d373d20af28b9d8e2cc0d0acf2334fcff457e47"
modules:
- "intel-mkl/2020.2.254"
- "python3/3.8.5"
- "netcdf/4.7.4p"
- "openmpi/4.1.2"
- "intel-compiler/2021.4.0"
keywords: [SIS2, symmetric, FMS2]
flags:
fflags: "-fno-alias -auto -safe-cray-ptr -ftz -assume byterecl -i4 -r8 -nowarn -sox -g -O2 -debug minimal -fp-model precise -qoverride-limits -xHost"
cflags: "-D__IFC -sox -g"
With the above information, one could then compile using the mom6-ninja-nci workflow written by @angus-g -- see here.
More information: - https://github.com/angus-g/mom6-ninja-nci - https://github.com/COSIMA/access-om2/wiki/Developers-guide (possibly out of date) - http://github.com/cosima/access-om3 (deprecated!)
How to find what diagnostics are available in MOM6
Presenter: @dougiesquire.
If one goes to to a run directory on Gadi, then you can look at available_diags.000000. This file tells you all the available diagnostics. For each of the disagnostics it lists, it tells you:
- which diagnostics are
[used]or[Unused]; - the units;
- description of the variable;
- name of the variable;
- which grid points the diagnostic is output on.
Here's an example:
"KHMEKE_v" [Unused]
! modules: {ocean_model,ocean_model_d2}
! dimensions: xh, yq
! long_name: Meridional diffusivity of MEKE
! units: m2 s-1
! cell_methods: xh:mean yq:point
If you are using an access-om3-configs configuration, we keep up to date versions of that file alongside our configurations, here. So the available_diags.000000 file is viewed here for the release-MC_25km_jra_iaf configuration.
How to add a diagnostic
One can simply add lines to the diag_table file (example). This is not the suggested workflow, we suggest editing this yaml file: https://github.com/ACCESS-NRI/access-om3-configs/blob/dev-MC_25km_jra_iaf%2Bwombatlite/diag_table_source.yaml
This yaml file is used by make_diag_table.py (latest version: https://github.com/COSIMA/make_diag_table) to create a diag_table file specifying MOM diagnostics.
Further information on the MOM diag_table format is here:
- https://github.com/mom-ocean/MOM5/blob/master/src/shared/diag_manager/diag_table.F90
- https://mom6.readthedocs.io/en/main/api/generated/pages/Diagnostics.html
- https://www.youtube.com/watch?v=D_J8eg3G80o
How to find how and where a diagnostic is calculated
Suppose one wants to know how a field is calculated. Taking our example above (and making sure we are looking at the correct mom6 code base for our configuration) we can search the mom6 code base for the long_name, namely Meridional diffusivity of MEKE, i.e. here.
So the relevant block is
CS%id_KhMEKE_v = register_diag_field('ocean_model', 'KHMEKE_v', diag%axesCv1, Time, &
'Meridional diffusivity of MEKE', 'm2 s-1', conversion=US%L_to_m**2*US%s_to_T)
Helps us find the actual array that is being output:
if (CS%id_KhMEKE_v>0) call post_data(CS%id_KhMEKE_v, Kh_v, CS%diag)
Kh_v.
Note that changing CICE outputs involves modifying the relevant namelist: ice_in (here is an example).
Overview of MOM6 configuration (input files etc)
Presenters: @aekiss @claireyung
Configuration and input files when running MOM6 "standalone" (FMS coupler). Actually, running MOM6 in an idealised way only requires 3 input files. Here is an example taken from Claire's IS-PG-MOM6 repository:
MOM_input-- parameter settings (name defined in 'MOM_input').diag_table-- diagnostics.input.nml-- run settings (calendar, MOM_input names etc).
As this is an idealised test case, much of the configuration (forcing, geometry etc) are defined in common/MOM_input. MOM_input file typically contain only the non-default values that are needed. A full list of parameters be found in the corresponding MOM_parameter_doc.all file (example) which is generated by the model at run-time. In Claire's case, she builds "perturbations" of the base case ("common") by modifying these files; in icemount-layer-LSPR (here) you can see the symlinks to the MOM_input, diag_table, input.nml files discussed above. Except that now there is a MOM_override file (here) which overrides any settings in MOM_input.
MOM6 also has an in-built sea-ice model SIS2. An example configuratin is here. You can see that there are more configuration files (additional to MOM_input, diag_table, input.nml):
field_table;SIS_inputSIS2 input files;data_tableforcing files if using FMS coupler
ACCESS-NRI ACCESS-OM3 uses a different coupler (NUOPC) compared to the above two examples. It also uses a different "standalone" sea-ice model CICE. So whilst the MOM elements discussed above remain the same. Additional files are required for NUOPC to couple the components.
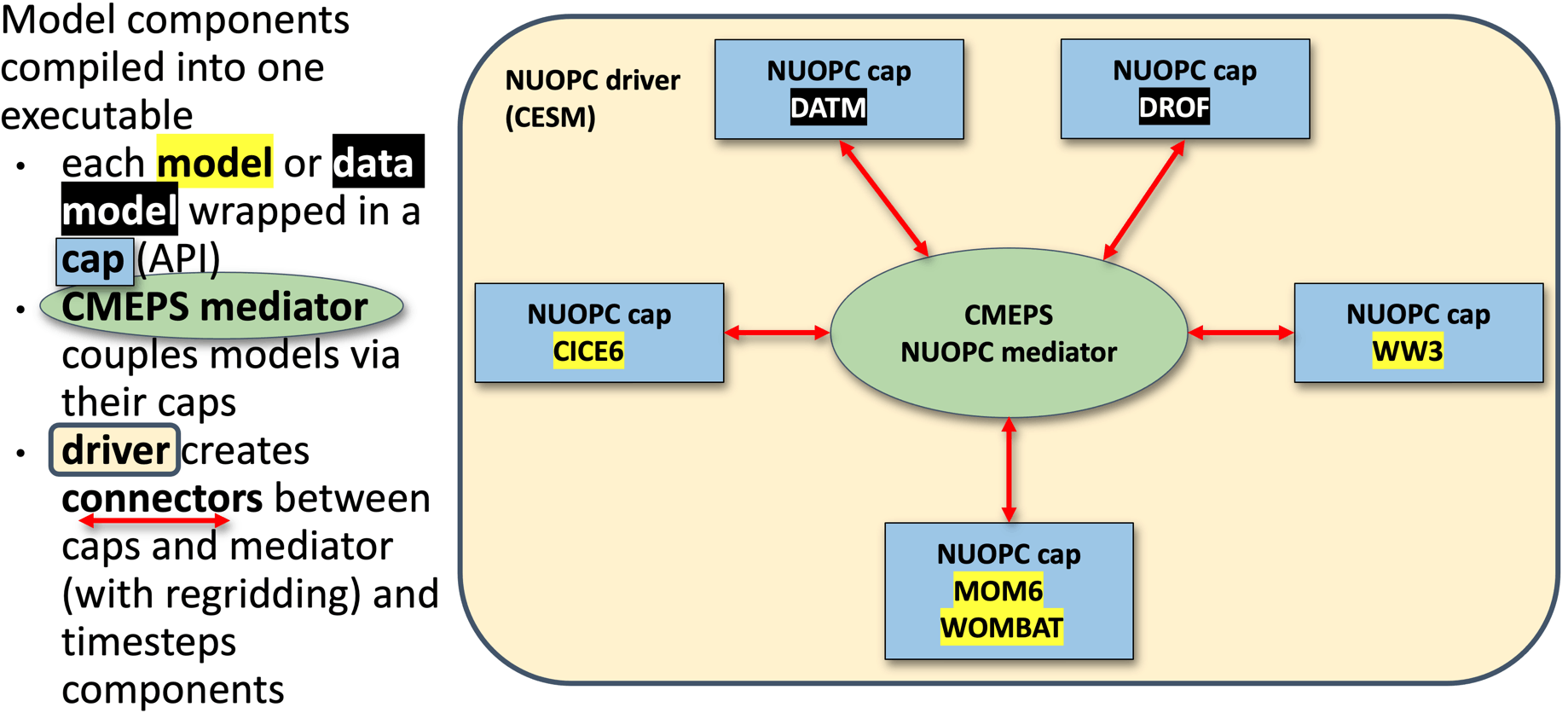 source: https://access-om3-configs.access-hive.org.au/infrastructure/Architecture/
source: https://access-om3-configs.access-hive.org.au/infrastructure/Architecture/
There's a description of the files found in a configuration here: https://access-om3-configs.access-hive.org.au/configurations/Overview/
OM3 configurations are stored here: https://github.com/acCESS-nri/access-om3-configs
Configurations that have a {dev|release}- prefix are the ones to focus on. Briefly, the configurations branches are named with the following
{dev|release}-{MODEL_COMPONENTS}_{nominal_resolution}km_{forcing_data}_{forcing_method}[+{modifier}]
Further details here.
Users are welcome to fork the access-nri configuration repository and share what changes/additions they make to configurations. ACCESS-NRI is also interested in helping support users share configurations (process outlined here). We are also keeping a list of key experiments used for development here.
Handy resources:
- ACCESS-NRI's OM3 configurations live here;
- ACCESS-NRI ACCESS-OM3 configuration files explanation;
- MOM6 runtime parameters format (input.nml, MOM_input);
- diag_table ;
- Another explanation of config files from MOM6 regional;
- AOS MOM6 tutorial 2022: Running and controlling MOM6 (e.g. ~15 minutes)